List of 5
$ ELA : tibble [3,627 × 27] (S3: tbl_df/tbl/data.frame)
..$ ssid : num [1:3627] 9466908 7683685 9025693 10099824 18886078 ...
..$ grade : int [1:3627] 11 11 11 11 11 11 11 11 11 11 ...
..$ year : int [1:3627] 11 11 11 11 11 11 11 11 11 11 ...
..$ content : chr [1:3627] "ELA" "ELA" "ELA" "ELA" ...
..$ testeventid : num [1:3627] 148933 147875 143699 143962 150680 ...
..$ asmtprmrydsbltycd : num [1:3627] 0 10 40 82 10 80 50 10 50 82 ...
..$ asmtscndrydsbltycd : num [1:3627] 0 0 20 0 0 80 0 0 0 0 ...
..$ Entry : num [1:3627] 123 88 105 153 437 307 305 42 59 304 ...
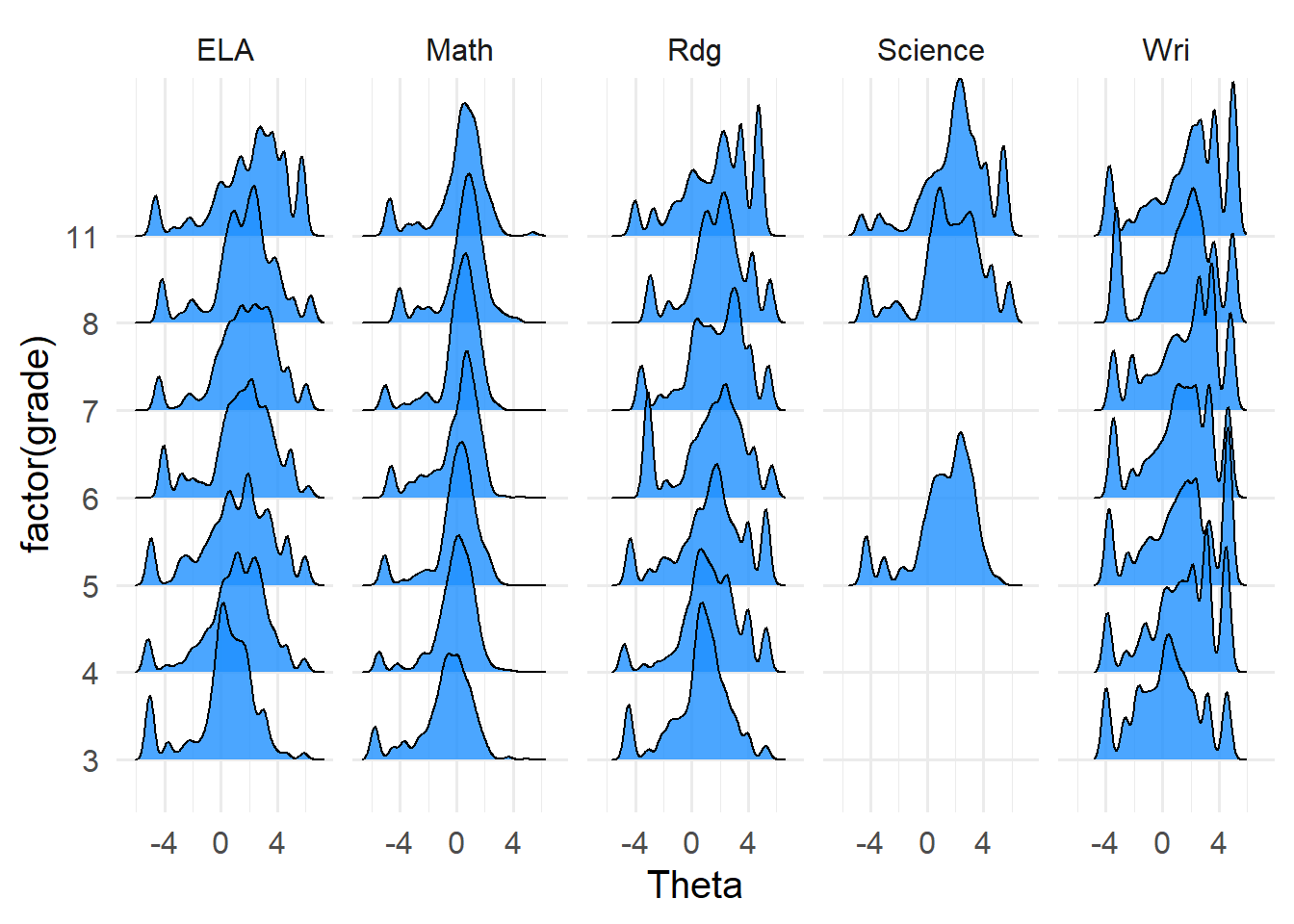
..$ Theta : num [1:3627] 1.27 1.55 3.28 4.48 2.67 ...
..$ Status : num [1:3627] 1 1 1 1 1 0 1 1 1 0 ...
..$ Count : num [1:3627] 36 36 36 36 36 36 36 36 36 36 ...
..$ RawScore : num [1:3627] 23 25 33 35 31 36 34 18 3 36 ...
..$ SE : num [1:3627] 0.371 0.385 0.619 1.023 0.501 ...
..$ Infit : num [1:3627] 0.93 0.95 0.9 0.93 0.92 1 1.06 1.55 0.85 1 ...
..$ Infit_Z : num [1:3627] -0.34 -0.37 -0.04 0.23 -0.18 0 0.31 2.56 0.06 0 ...
..$ Outfit : num [1:3627] 0.82 0.81 1.63 0.35 0.88 1 0.86 1.74 0.17 1 ...
..$ Outfit_Z : num [1:3627] -0.62 -0.56 1.03 -0.16 -0.12 0 0.17 3.31 -0.37 0 ...
..$ Displacement : num [1:3627] 0.0018 0.0019 0.0022 0.0023 0.0021 0.0024 0.0022 0.0017 0.0009 0.0024 ...
..$ PointMeasureCorr : num [1:3627] 0.42 0.42 0.3 0.27 0.31 0 0.14 -0.12 0.32 0 ...
..$ Weight : num [1:3627] 1 1 1 1 1 1 1 1 1 1 ...
..$ ObservMatch : num [1:3627] 75 80.6 91.7 97.2 86.1 100 94.4 50 97.2 100 ...
..$ ExpectMatch : num [1:3627] 68.3 72 91.7 97.2 86.1 100 94.4 65 97.2 100 ...
..$ PointMeasureExpected: num [1:3627] 0.35 0.33 0.2 0.12 0.25 0 0.17 0.38 0.17 0 ...
..$ RMSR : num [1:3627] 0.42 0.39 0.26 0.16 0.31 0 0.23 0.51 0.14 0 ...
..$ WMLE : num [1:3627] 1.25 1.68 3.13 4 2.59 ...
..$ primary : Factor w/ 12 levels "0","10","20",..: 1 2 4 11 2 10 6 2 6 11 ...
..$ secondary : Factor w/ 12 levels "0","10","20",..: 1 1 3 1 1 10 1 1 1 1 ...
$ Math : tibble [3,629 × 27] (S3: tbl_df/tbl/data.frame)
..$ ssid : num [1:3629] 10129634 10926496 10616063 10139443 8887381 ...
..$ grade : int [1:3629] 11 11 11 11 11 11 11 11 11 11 ...
..$ year : int [1:3629] 11 11 11 11 11 11 11 11 11 11 ...
..$ content : chr [1:3629] "Math" "Math" "Math" "Math" ...
..$ testeventid : num [1:3629] 147564 151249 146976 151229 147676 ...
..$ asmtprmrydsbltycd : num [1:3629] 10 80 10 10 80 82 10 10 40 82 ...
..$ asmtscndrydsbltycd : num [1:3629] 0 90 0 0 10 80 50 0 50 0 ...
..$ Entry : num [1:3629] 161 344 321 167 101 131 141 357 14 367 ...
..$ Theta : num [1:3629] 0.514 1.965 0.899 1.965 -4.724 ...
..$ Status : num [1:3629] 1 1 1 1 -1 1 1 1 1 1 ...
..$ Count : num [1:3629] 36 36 36 36 36 36 36 36 36 36 ...
..$ RawScore : num [1:3629] 19 29 22 29 0 14 18 25 19 22 ...
..$ SE : num [1:3629] 0.356 0.436 0.362 0.436 1.839 ...
..$ Infit : num [1:3629] 0.97 1.03 0.99 1 1 1.01 0.99 0.85 0.96 1.06 ...
..$ Infit_Z : num [1:3629] -0.18 0.26 -0.09 0.1 0 0.13 -0.06 -0.81 -0.39 0.5 ...
..$ Outfit : num [1:3629] 0.97 1.03 1.01 0.87 1 0.93 1 0.77 0.97 0.99 ...
..$ Outfit_Z : num [1:3629] -0.17 0.23 0.17 -0.46 0 -0.03 0.03 -0.87 -0.19 0.07 ...
..$ Displacement : num [1:3629] 5e-04 5e-04 5e-04 5e-04 -7e-04 4e-04 5e-04 5e-04 5e-04 5e-04 ...
..$ PointMeasureCorr : num [1:3629] 0.38 0.28 0.34 0.29 0 0.34 0.38 0.45 0.38 0.27 ...
..$ Weight : num [1:3629] 1 1 1 1 1 1 1 1 1 1 ...
..$ ObservMatch : num [1:3629] 66.7 80.6 66.7 80.6 100 61.1 69.4 75 63.9 61.1 ...
..$ ExpectMatch : num [1:3629] 64.4 80.5 66 80.5 100 68.3 64.6 70.3 64.4 66 ...
..$ PointMeasureExpected: num [1:3629] 0.35 0.25 0.33 0.25 0 0.35 0.35 0.3 0.35 0.33 ...
..$ RMSR : num [1:3629] 0.46 0.39 0.45 0.39 0 0.46 0.46 0.4 0.46 0.48 ...
..$ WMLE : num [1:3629] 0.512 1.915 0.888 1.915 -4.724 ...
..$ primary : Factor w/ 12 levels "0","10","20",..: 2 10 2 2 10 11 2 2 4 11 ...
..$ secondary : Factor w/ 12 levels "0","10","20",..: 1 12 1 1 2 10 6 1 6 1 ...
$ Rdg : tibble [3,627 × 27] (S3: tbl_df/tbl/data.frame)
..$ ssid : num [1:3627] 18631185 18342736 10897771 7663935 6709613 ...
..$ grade : int [1:3627] 11 11 11 11 11 11 11 11 11 11 ...
..$ year : int [1:3627] 11 11 11 11 11 11 11 11 11 11 ...
..$ content : chr [1:3627] "Rdg" "Rdg" "Rdg" "Rdg" ...
..$ testeventid : num [1:3627] 145641 147717 145196 149014 146545 ...
..$ asmtprmrydsbltycd : num [1:3627] 10 90 10 80 0 10 10 80 82 82 ...
..$ asmtscndrydsbltycd : num [1:3627] 0 0 0 10 0 82 0 70 0 0 ...
..$ Entry : num [1:3627] 444 444 343 74 3 77 181 261 297 182 ...
..$ Theta : num [1:3627] 2.24 3.48 2.72 -2.72 0.12 -1.39 -0.67 4.73 4.73 4.73 ...
..$ Status : num [1:3627] 1 1 1 1 1 1 1 0 0 0 ...
..$ Count : num [1:3627] 19 19 19 19 19 19 19 19 19 19 ...
..$ RawScore : num [1:3627] 16 18 17 1 8 3 5 19 19 19 ...
..$ SE : num [1:3627] 0.64 1.03 0.76 1.06 0.49 0.66 0.55 1.84 1.84 1.84 ...
..$ Infit : num [1:3627] 1.07 1.02 1.1 1 0.98 0.73 1.08 1 1 1 ...
..$ Infit_Z : num [1:3627] 0.29 0.31 0.53 0.29 -0.11 -0.54 0.37 0 0 0 ...
..$ Outfit : num [1:3627] 1.17 0.94 1.22 0.76 0.95 0.62 1.09 1 1 1 ...
..$ Outfit_Z : num [1:3627] 0.45 0.32 0.71 0 -0.22 -0.62 0.42 0 0 0 ...
..$ Displacement : num [1:3627] 0 0 0 0 0 0 0 0 0 0 ...
..$ PointMeasureCorr : num [1:3627] 0.01 0.08 0.05 0 0.32 0.68 0.2 0 0 0 ...
..$ Weight : num [1:3627] 1 1 1 1 1 1 1 1 1 1 ...
..$ ObservMatch : num [1:3627] 84.2 94.7 89.5 94.7 63.2 84.2 78.9 100 100 100 ...
..$ ExpectMatch : num [1:3627] 84.2 94.7 89.5 94.7 64.2 85.1 76.3 100 100 100 ...
..$ PointMeasureExpected: num [1:3627] 0.17 0.1 0.14 0.24 0.3 0.31 0.32 0 0 0 ...
..$ RMSR : num [1:3627] 0.38 0.22 0.31 0.22 0.47 0.34 0.36 0 0 0 ...
..$ WMLE : num [1:3627] 2.11 3.02 2.5 -2.27 0.14 -0.9 -0.6 4.73 4.73 4.73 ...
..$ primary : Factor w/ 12 levels "0","10","20",..: 2 12 2 10 1 2 2 10 11 11 ...
..$ secondary : Factor w/ 12 levels "0","10","20",..: 1 1 1 2 1 11 1 8 1 1 ...
$ Science: tibble [1,435 × 27] (S3: tbl_df/tbl/data.frame)
..$ ssid : num [1:1435] 7617607 7642717 10341706 9811494 10469745 ...
..$ grade : int [1:1435] 11 11 11 11 11 11 11 11 11 11 ...
..$ year : int [1:1435] 11 11 11 11 11 11 11 11 11 11 ...
..$ content : chr [1:1435] "Science" "Science" "Science" "Science" ...
..$ testeventid : num [1:1435] 148838 144146 149634 146456 144426 ...
..$ asmtprmrydsbltycd : num [1:1435] 10 82 80 10 90 10 10 80 80 10 ...
..$ asmtscndrydsbltycd : num [1:1435] 0 80 20 0 50 80 70 0 50 0 ...
..$ Entry : num [1:1435] 28 43 274 118 291 91 58 130 297 337 ...
..$ Theta : num [1:1435] 0.875 2.647 4.17 5.404 5.404 ...
..$ Status : num [1:1435] 1 1 1 0 0 1 1 1 1 1 ...
..$ Count : num [1:1435] 36 36 36 36 36 36 36 36 36 36 ...
..$ RawScore : num [1:1435] 22 32 35 36 36 26 2 34 34 28 ...
..$ SE : num [1:1435] 0.361 0.544 1.021 1.835 1.835 ...
..$ Infit : num [1:1435] 1.3 0.94 1.05 1.04 0.99 0.9 0.97 0.99 0.98 0.9 ...
..$ Infit_Z : num [1:1435] 2.35 0.02 0.37 0.35 0 -0.62 0.17 0.15 0.16 -0.52 ...
..$ Outfit : num [1:1435] 1.22 0.76 1.5 1.2 0.48 0.85 0.69 1.57 0.67 0.8 ...
..$ Outfit_Z : num [1:1435] 1.87 -0.27 0.91 0.6 -0.16 -0.87 -0.41 0.74 0.01 -0.76 ...
..$ Displacement : num [1:1435] 0.0011 0.0016 0.0016 0.0027 0.0027 0.0013 0.0007 0.0016 0.0016 0.0014 ...
..$ PointMeasureCorr : num [1:1435] 0.07 0.3 -0.07 0 0.29 0.46 0.28 0.05 0.22 0.45 ...
..$ Weight : num [1:1435] 1 1 1 1 1 1 1 1 1 1 ...
..$ ObservMatch : num [1:1435] 55.6 88.9 97.2 100 100 75 94.4 94.4 94.4 80.6 ...
..$ ExpectMatch : num [1:1435] 66.1 88.9 97.2 100 100 73.2 94.4 94.4 94.4 77.9 ...
..$ PointMeasureExpected: num [1:1435] 0.32 0.22 0.12 0 0 0.3 0.15 0.16 0.16 0.28 ...
..$ RMSR : num [1:1435] 0.52 0.3 0.17 0 0 0.39 0.22 0.22 0.22 0.36 ...
..$ WMLE : num [1:1435] 0.863 2.543 3.692 5.404 5.404 ...
..$ primary : Factor w/ 12 levels "0","10","20",..: 2 11 10 2 12 2 2 10 10 2 ...
..$ secondary : Factor w/ 12 levels "0","10","20",..: 1 10 3 1 6 10 8 1 6 1 ...
$ Wri : tibble [3,627 × 27] (S3: tbl_df/tbl/data.frame)
..$ ssid : num [1:3627] 7653093 7640498 7650957 9305807 18060455 ...
..$ grade : int [1:3627] 11 11 11 11 11 11 11 11 11 11 ...
..$ year : int [1:3627] 11 11 11 11 11 11 11 11 11 11 ...
..$ content : chr [1:3627] "Wri" "Wri" "Wri" "Wri" ...
..$ testeventid : num [1:3627] 148893 148868 144397 148988 144865 ...
..$ asmtprmrydsbltycd : num [1:3627] 10 10 90 0 82 10 80 0 10 82 ...
..$ asmtscndrydsbltycd : num [1:3627] 0 0 80 0 0 0 50 0 0 10 ...
..$ Entry : num [1:3627] 61 48 63 107 443 214 267 344 423 27 ...
..$ Theta : num [1:3627] 1.67 2.17 -1.56 5.01 2.17 0.38 0.79 5.01 -0.48 2.78 ...
..$ Status : num [1:3627] 1 1 1 0 1 1 1 0 1 1 ...
..$ Count : num [1:3627] 13 13 13 13 13 13 13 13 13 13 ...
..$ RawScore : num [1:3627] 9 10 2 13 10 6 7 13 4 11 ...
..$ SE : num [1:3627] 0.69 0.74 0.83 1.87 0.74 0.64 0.65 1.87 0.68 0.84 ...
..$ Infit : num [1:3627] 0.52 0.65 0.61 1 1.01 0.46 0.47 1 0.9 0.88 ...
..$ Infit_Z : num [1:3627] -2.07 -1.02 -0.79 0 0.24 -2.3 -2.2 0 -0.16 -0.21 ...
..$ Outfit : num [1:3627] 0.44 0.52 0.33 1 0.74 0.41 0.4 1 1.32 0.54 ...
..$ Outfit_Z : num [1:3627] -1.65 -0.81 -0.47 0 -0.06 -1.88 -1.85 0 -0.14 -0.16 ...
..$ Displacement : num [1:3627] 0 0 0 0 0 0 0 0 0 0 ...
..$ PointMeasureCorr : num [1:3627] 0.86 0.73 0.62 0 0.51 0.87 0.87 0 0.56 0.56 ...
..$ Weight : num [1:3627] 1 1 1 1 1 1 1 1 1 1 ...
..$ ObservMatch : num [1:3627] 100 92.3 84.6 100 76.9 100 100 100 92.3 84.6 ...
..$ ExpectMatch : num [1:3627] 75.5 79.7 84.6 100 79.7 72.9 73 100 74.5 84.6 ...
..$ PointMeasureExpected: num [1:3627] 0.48 0.44 0.34 0 0.44 0.5 0.51 0 0.45 0.38 ...
..$ RMSR : num [1:3627] 0.31 0.34 0.26 0 0.38 0.29 0.29 0 0.37 0.31 ...
..$ WMLE : num [1:3627] 1.61 2.08 -1.38 5.01 2.08 0.38 0.78 5.01 -0.43 2.61 ...
..$ primary : Factor w/ 12 levels "0","10","20",..: 2 2 12 1 11 2 10 1 2 11 ...
..$ secondary : Factor w/ 12 levels "0","10","20",..: 1 1 10 1 1 1 6 1 1 2 ...